MELANOCORTIN RECEPTORS
Melanocortin receptors are G protein-coupled receptors (GPCRs) for melanocortins, which compose a key modulatory system for multiple physiological roles. Five subtypes of melanocortin receptors (MC1–MC5) were identified during the early 1990s. These receptors are one of few GPCRs in that they are regulated by both endogenous agonists (proopiomelanocortin derivatives) and inverse agonists (agouti-related protein). Our research group’s main focus among melanocortin receptors is on studying ligand binding process and activation of MC1 and MC3-MC5 receptors.
MC1 receptors are involved in regulation of skin and hair color in mammalians and in development of melanoma, where they are overexpressed. To characterize the pharmacological properties of melanocortin ligands we have tested the BacMam-Epac-cAMP sensor on melanocortin receptors on mammalian cells. The sensor distinguishes full and partial agonists and antagonists (Mazina et al. 2012).
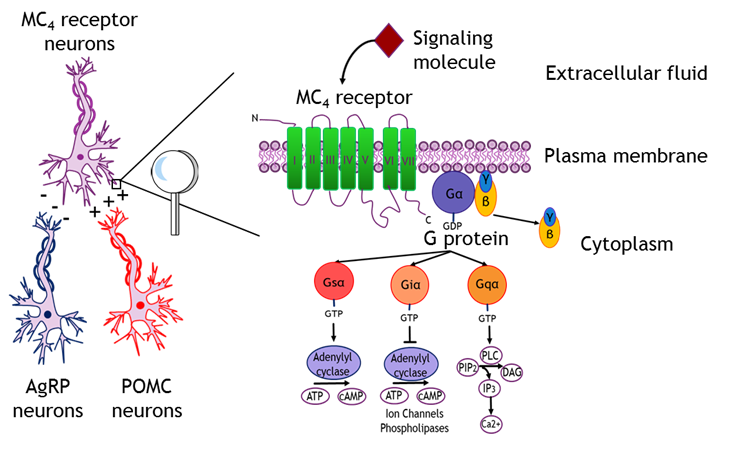
MC4 receptors are known as a regulator of eating behavior and body weight, as a modulator of erectile function and sexual behavior, and as a potential target for treatment of depression and anxiety disorders (Figure 1.).
MC1 receptors are involved in regulation of skin and hair color in mammalians and in development of melanoma, where they are overexpressed. To characterize the pharmacological properties of melanocortin ligands we have tested the BacMam-Epac-cAMP sensor on melanocortin receptors on mammalian cells. The sensor distinguishes full and partial agonists and antagonists (Mazina et al. 2012).
MC4 receptors are known as a regulator of eating behavior and body weight, as a modulator of erectile function and sexual behavior, and as a potential target for treatment of depression and anxiety disorders (Figure 1.).
Figure 1. Melanocortin 4 (MC4) receptors are G protein-coupled receptors that are regulated by both endogenous agonists and inverse agonists released from proopiomelanocortin (POMC) and agouti-related protein (AgRP) neurons, respectively. Upon activation these G protein-coupled receptors may be associated with three types of G proteins: Gsα, Giα or Gqα.
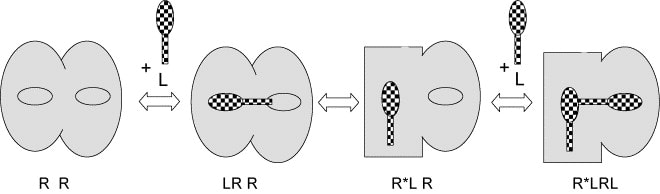
The ligand binding to these G protein-coupled receptors is subject to considerable complexity. Melanocortin receptor activation model was established in 2006 (Figure 2.): allosteric model with two tandem binding sites (Kopanchuk et al. 2006). This model was established based on binding data of radioligand binding assay on Sf9 cell membranes. In addition to complex dynamic regulation, other effector molecules like accessory proteins have also been found to potentially influence the binding of melanocortin MC4 receptors. Sf9/baculovirus expression system can be used for the coexpression of receptors and their accessory proteins.
Figure 2. Schematic illustration of the allosteric two tandem binding-site model of ligand binding to an MC4 receptor dimer and the conformational transformations associated therewith.
To study this complex dynamic regulation model further we moved from radioactivity to fluorescence methods to be able to get more information on both time and space. One of the promising fluorescence-based methods that allows continuous online monitoring of receptor-ligand interactions is fluorescence anisotropy (FA). The studies of receptors on a molecular level require specific and high-affinity reporter ligands. With fluorescence anisotropy measurements we have characterized four fluorescent ligands: Cy3B-NDP-α-MSH, TAMRA-α-MSH, UTBC101 and UTBC102 (Veiksina et al. 2010, Veiksina et al. 2014, Link et al. 2017). These reporter ligands exhibit nanomolar affinity toward melanocortin MC4 receptors but have relatively different kinetic properties as well as pharmacological properties. Cy3B-NDP-α-MSH was first used in baculovirus-display system, which possesses superior signal-to-noise ratio compared to the previous cell membrane assays (Veiksina et al. 2014) and (Veiksina et al. 2015).
MC4 receptor ligands Cy3B-NDP, UTBC101 and UTBC102 in baculovirus-display system can be used outside pseudo-first order system and analysed using global analysis with differential equations. With global analysis we can account for many different effects existing in real situation, e.g., ligand bleaching and degradation in time (Veiksina et al. 2014, Link et al. 2017). As these experiments produce large amounts of data there is a need for software that enables easy data manipulation. Approriate open source software developed in our laboratory can be found from here.
Reporter ligands are used for the characterization of affinity and potency of unlabelled ligands. We have demostrated on the MC4 receptor-ligand system that the equilibrium in the competition experiment depends on the kinetic properties of both reporter and unlabelled ligand. Ignoring this kinetic aspect may lead to dramatic over- or underestimation of an unlabelled ligand’s binding affinity (Kopanchuk et al. 2006, Veiksina et al. 2010, Link et al. 2017, Rinken et al. 2016). Kinetic parameters of unlabelled ligands can be characterised from the numerical fitting of displacement curves in time (Rinken et al. TIPS, in press).
MC4 receptor ligands Cy3B-NDP, UTBC101 and UTBC102 in baculovirus-display system can be used outside pseudo-first order system and analysed using global analysis with differential equations. With global analysis we can account for many different effects existing in real situation, e.g., ligand bleaching and degradation in time (Veiksina et al. 2014, Link et al. 2017). As these experiments produce large amounts of data there is a need for software that enables easy data manipulation. Approriate open source software developed in our laboratory can be found from here.
Reporter ligands are used for the characterization of affinity and potency of unlabelled ligands. We have demostrated on the MC4 receptor-ligand system that the equilibrium in the competition experiment depends on the kinetic properties of both reporter and unlabelled ligand. Ignoring this kinetic aspect may lead to dramatic over- or underestimation of an unlabelled ligand’s binding affinity (Kopanchuk et al. 2006, Veiksina et al. 2010, Link et al. 2017, Rinken et al. 2016). Kinetic parameters of unlabelled ligands can be characterised from the numerical fitting of displacement curves in time (Rinken et al. TIPS, in press).